 
The host instead uses a synthetic amino acid, selenomethionine, in which methionene's sulfur atom has been replaced by selenium. Molecular replacement is used extensively in labs that solve the crystal structures of several mutants of a given protein.Īnomalous x-ray scattering relies on protein production in a host that is incapable of producing the amino acid methionine. The technique of molecular replacement uses the solved crystal structure of a homologous protein to provide a "seed" electron density map that can then be refined by a computer. The 1985 Nobel Prize in chemistry was awarded to Hauptman and Karle, who developed these methods. These methods are only viable for small (less than 1000 atoms) molecules and are not typically used in protein crystallography. Common techniques include:ĭirect methods use the Sayre equation to determine phases directly from the diffraction data. Crystallographers employ computational methods to capture the x-ray scattering pattern (pictured at right) and infer the three-dimensional positions of atoms in a molecule.Ĭrystallographers use several methods for recovering phase information from diffraction data. However, x-rays are difficult to focus in a manner analogous to the way a lens focuses visible light. X-rays, where $100 pm < \(\lambda\) < 10,000 pm$, are the perfect size to diffract around atoms (32-225 pm), bonds (74-267 pm), and molecules (100 pm to hundreds of Angstroms). Molecules such as proteins are much smaller than microscopic structures like cells, and, as such, require that a shorter wavelength of radiation be used during diffraction. In a light microscope, the subject is irradiated with visible light (400 nm < \( \lambda \) < 700 nm$), which is diffracted by a lens onto the retina, producing a macroscopic image of a microscopic object. An x-ray diffractometer works in a similar manner to a light microscope. Once crystals of suitable size and composition are obtained, it is necessary to bombard the crystal with x-rays and observe the diffraction pattern. Patience, and to some extent, luck, determine the sucess or failure of the crystallization of any particular protein. It is often said that this part of crystallography is more of an art than a science, and indeed there is little theoretical guidance available to the crystallographer who wishes to crystallize a new protein. 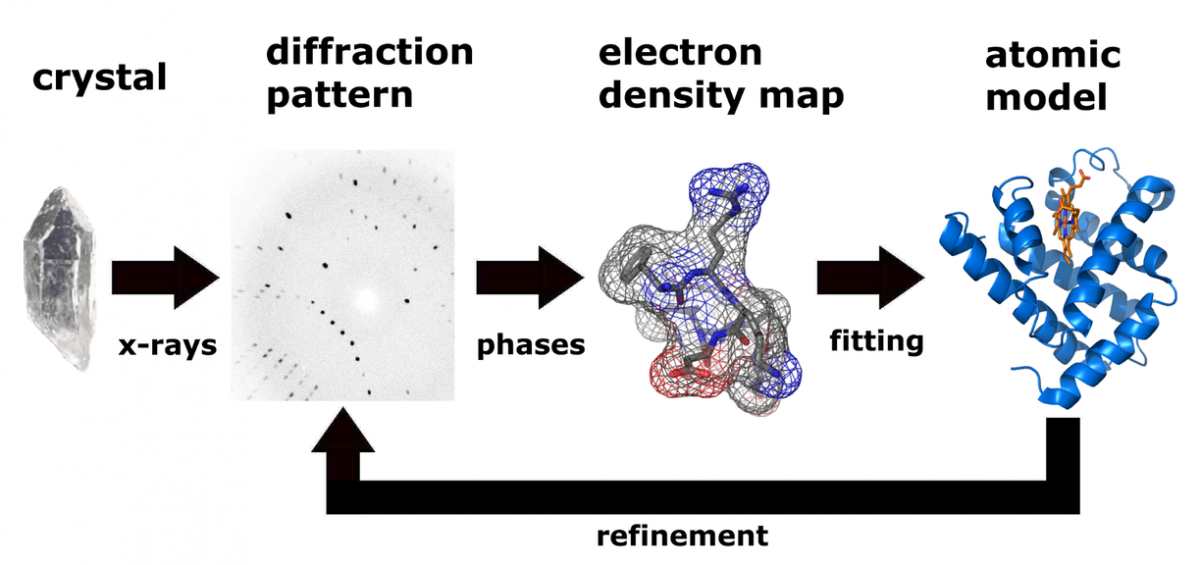
If this process occurs at just the right rate, the protein precipitates out of solution into an ordered lattice structure: a protein crystal. Concomitantly, the concentration of protein increases. The drop is hypotonic to the precipitant and slowly equilibrates to the concentration of the large volume of precipitant. This drop is separated by air from a large volume of precipitant solution. In this technique, a small quantity (typically a microliter) of concentrated protein solution is mixed with an equal volume of precipitant. Since we have no way of knowing a priori which set of conditions is right for obtaining crystals of a given protein, many different conditions are tried in parallel using a technique called drop diffusion. The concentrated protein solution obtained is then subjected to a wide variety of crystallization conditions. The PDB began with 13 structures in 1976 and has grown to the " single worldwide archive of structural data of biological macromolecules". The early 70s saw the birth of the Research Collaboratory for Structural Bioinformatics' Protein Data Bank. David Chilton Phillips solved the first structure of an enzyme, lysozyme, in 1965. The field has received numerous Nobel Prizes over the years, including in chemistry in 1964 to Dorothy Crowfoot Hodgkin, who solved the structures of the small molecules cholesterol, penicillin, and vitamin B12, and in chemistry in 1962 to Max Perutz and John Cowdery Kendrew for their work on sperm whale myoglobin. Which relates an x-ray diffraction pattern with the three-dimensional structure of a crystal. Though crystal symmetry was explored in the late 1600s by Danish scientist Nicolas Steno, and continuing efforts by René Just Haüy and William Hallowes Miller in 1839 firmly established that a crystal is a ordered lattice, it wasn't until the discovery of x-rays in 1895 and the proof of their diffraction by Max von Laue in 1912 that crystallography as a science began.Īfter the use of x-ray crystallography to deduce the lattice structure of table salt in 1914, the father and son team of William Henry Bragg and William Lawrence Bragg shared the 1915 Nobel Prize in Physics for the development of Bragg's law, 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
